iso-hiEBS, hiEBS clone 8
MLi002-A-1
General
Cell Line |
|
| hPSCreg name | MLi002-A-1 |
| Cite as: | MLi002-A-1 |
| Alternative name(s) |
iso-hiEBS, hiEBS clone 8
|
| Cell line type | Human induced pluripotent stem cell (hiPSC) |
| Similar lines | No similar lines found. |
| Last update | 7th April 2026 |
| User feedback | |
Provider |
|
| Generator | Faculty of Medicine University of Ljubljana (ML) |
| Owner | Faculty of Medicine University of Ljubljana (ML) |
| Distributors | |
| Derivation country | Netherlands |
External Databases |
|
| BioSamples | SAMEA121967211 |
General Information |
|
| * Is the cell line readily obtainable for third parties? |
No |
| Subclone of | |
Genetic Modification
| Disease/phenotype related modifications |
|
Donor Information
General Donor Information |
|
| Sex | female |
| Ethnicity | Caucasian |
Phenotype and Disease related information (Donor) |
|
| Diseases | A disease was diagnosed.
|
External Databases (Donor) |
|
| BioSamples | SAMEA5340319 |
Ethics
Also have a look at the ethics information for the parental line
MLi002-A
.
| Is there an MTA available for the cell line? | No |
| For generation of the cell line, who was the supplier of any recombined DNA vectors or commercial kits used? | Integrated DNA Technologies |
| Are you aware of any constraints on the use or distribution of the cell line from the owner or any parties identified in the query above? | Yes |
| Constraints for use or distribution | Cells are not available to researchers anywhere in the world; there is controlled access to the genetic information associated with the cell line; it is permitted to store cells for an unlimited time; it is not permitted to use a cell line for clinical treatment or human applications. |
hIPSC Derivation
General |
|
|
The source cell information can be found in the parental cell line
MLi002-A.
|
|
| Passage number reprogrammed | 3 |
Reprogramming method |
|
| Vector type | Non-integrating |
| Vector | Sendai virus |
| Genes | |
| Is reprogramming vector detectable? |
Yes |
| Methods used |
PCR, RT-PCR, Sequencing
|
| Notes on reprogramming vector detection | CytoTune-iPS 2.0 Sendai Reprogramming Kit from Invitrogen. After 10 passages, elimination of the SeV vectors was confirmed in the parental MLi002-A cell line by RT-PCR. For CRISPR–Cas9 editing, passage 19 was used; therefore, the presence of SeV was not rechecked. |
Vector free reprogramming |
|
| Type of used vector free reprogramming factor(s) |
None
|
Other |
|
| Selection criteria for clones | Single cell-derived colonies (clones) were manually picked and screened for the corrected genotype using restriction analysis. |
| Derived under xeno-free conditions |
Yes |
| Derived under GMP? |
No |
| Available as clinical grade? |
No |
Culture Conditions
| Surface coating | Vitronectin |
| Feeder cells |
No |
| Passage method |
Enzyme-free cell dissociation
Gentle Cell Dissociation Reagent
|
| O2 Concentration | 20 % |
| CO2 Concentration | 5 % |
| Medium |
Other medium:
Base medium: StemFlex medium (Gibco), TeSR-AOF medium (StemCell Technologies)
Main protein source: Serum concentration: % |
| Has Rock inhibitor (Y27632) been used at passage previously with this cell line? | Yes |
| Has Rock inhibitor (Y27632) been used at cryo previously with this cell line? | No |
| Has Rock inhibitor (Y27632) been used at thaw previously with this cell line? | Yes |
Characterisation
Analysis of Undifferentiated Cells
| Marker | Expressed | Immunostaining | RT-PCR | Flow Cytometry | Enzymatic Assay | Expression Profiles |
| NANOG |
Yes |
|||||
| POU5F1 (OCT-4) |
Yes |
|||||
| SSEA-4 |
Yes |
Differentiation Potency
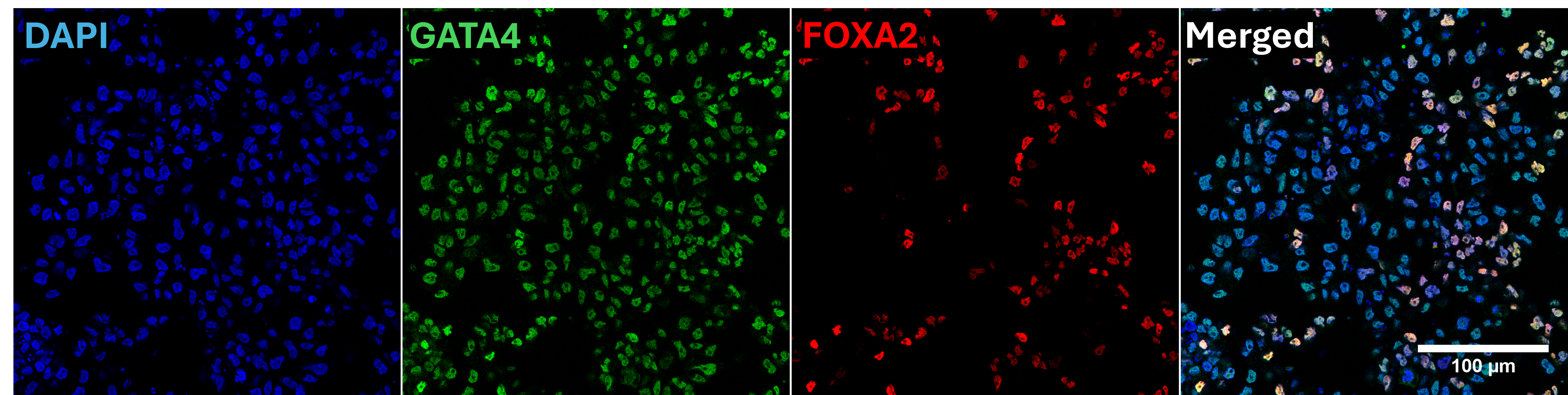
In vitro directed differentiation
| Marker | Expressed |
| GATA4 |
Yes |
| FOXA2 |
Yes |
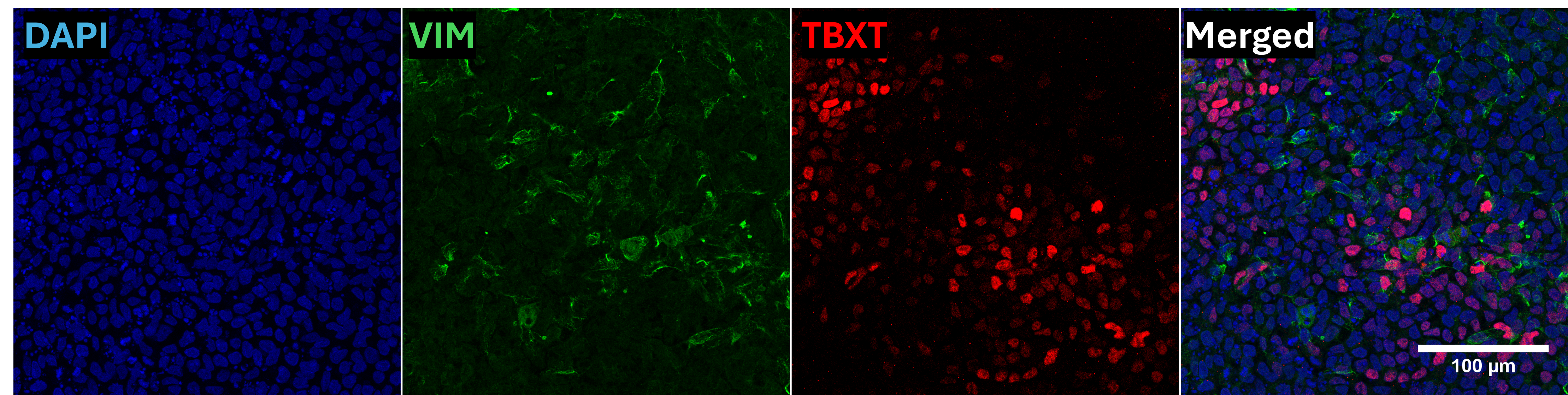
In vitro directed differentiation
| Marker | Expressed |
| VIM |
Yes |
| TBXT (Brachyury) |
Yes |
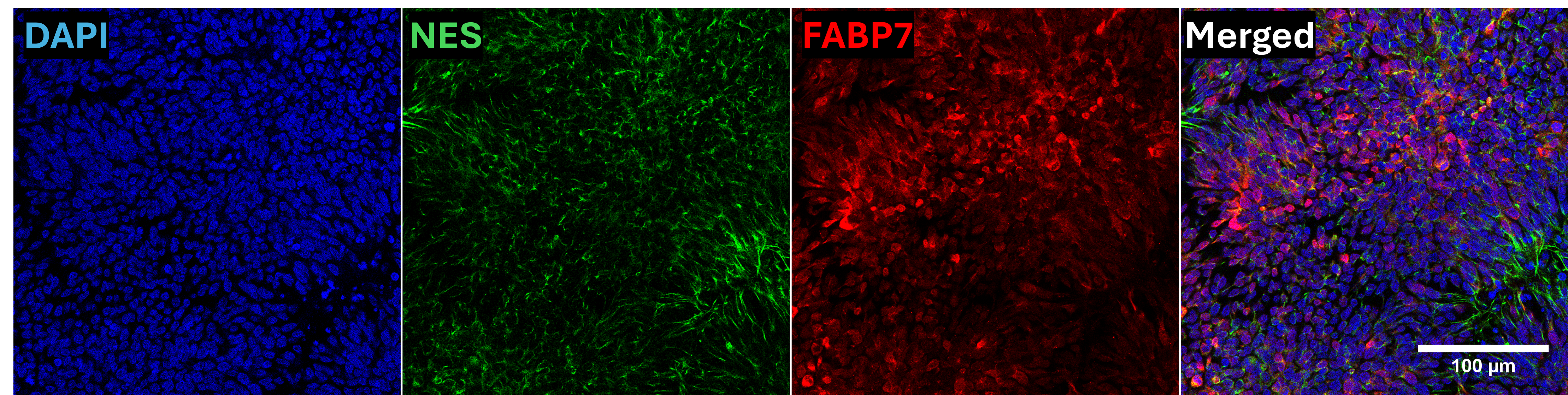
In vitro directed differentiation
| Marker | Expressed |
| FABP7 |
Yes |
| NES |
Yes |
Microbiology / Virus Screening |
|
| Mycoplasma | Negative |
Genotyping
Karyotyping (Cell Line) |
|
| Has the cell line karyotype been analysed? |
Yes
|
Other Genotyping (Cell Line) |
|


Login to share your feedback, experiences or results with the research community.